OpenGeneMed
Developed by: Alida Palmisano, Ming-Chung Li, Eric Polley, Yingdong Zhao*, Richard Simon*
OpenGeneMed is an informatics system to support clinical trial research.
OpenGeneMed is an open and customizable version of the GeneMed system (Zhao et al,
2015), a web-based interface developed for the Molecular Profiling based Assignment of
Cancer Therapy (NCI-MPACT) clinical trial coordinated by the NIH.
OpenGeneMed streamlines clinical trial management and it can be used by clinicians, lab
personnel, statisticians and other researchers as a communication hub. It automates the
annotation of the genomic variants identified by sequencing, classifies the actionable mutations
according to customizable rules, and facilitates quality control by the molecular
characterization lab in the review of variants.
OpenGeneMed collects baseline information of patients to determine eligibility for the panel of
treatment regimens available. The system generates patient reports containing detected genomic
alterations along with summarized information that can be used for treatment assignment by a
supervising treatment review team.
OpenGeneMed is distributed as a standalone virtual machine, ready for deployment and use from a
web browser. OpenGeneMed code is modular which allows customization of existing features and
addition of new modules to address the specific needs of different clinical trials and research
teams. Examples on how to customize the code are provided in the technical documentation
distributed with the virtual machine.
In summary, OpenGeneMed offers an initial set of features inspired by our experience with
GeneMed, a system that has been proven to be an efficient and successful informatics hub for
coordinating the application of next-generation sequencing in the NCI-MPACT trial.
*corresponding author
Acknowledgment: we thank Henry Rivera for a preliminary implementation of the
OpenGeneMed.
Screenshots from OpenGeneMed
Please refer to the User Manual for more details and full description of all the software features.
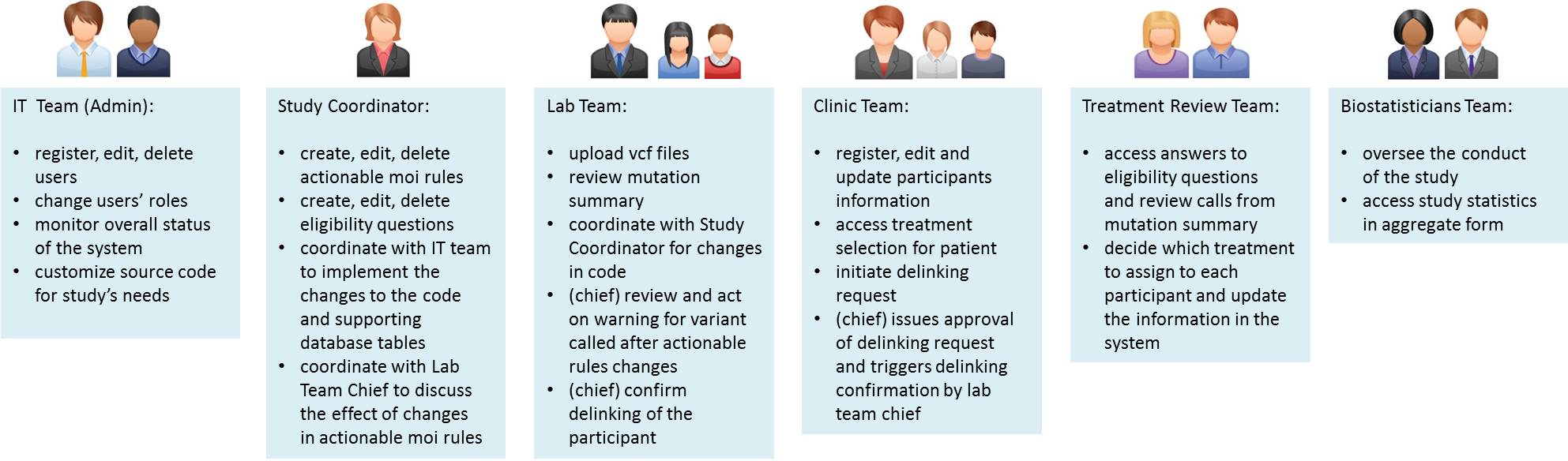
Groups available in the default setup
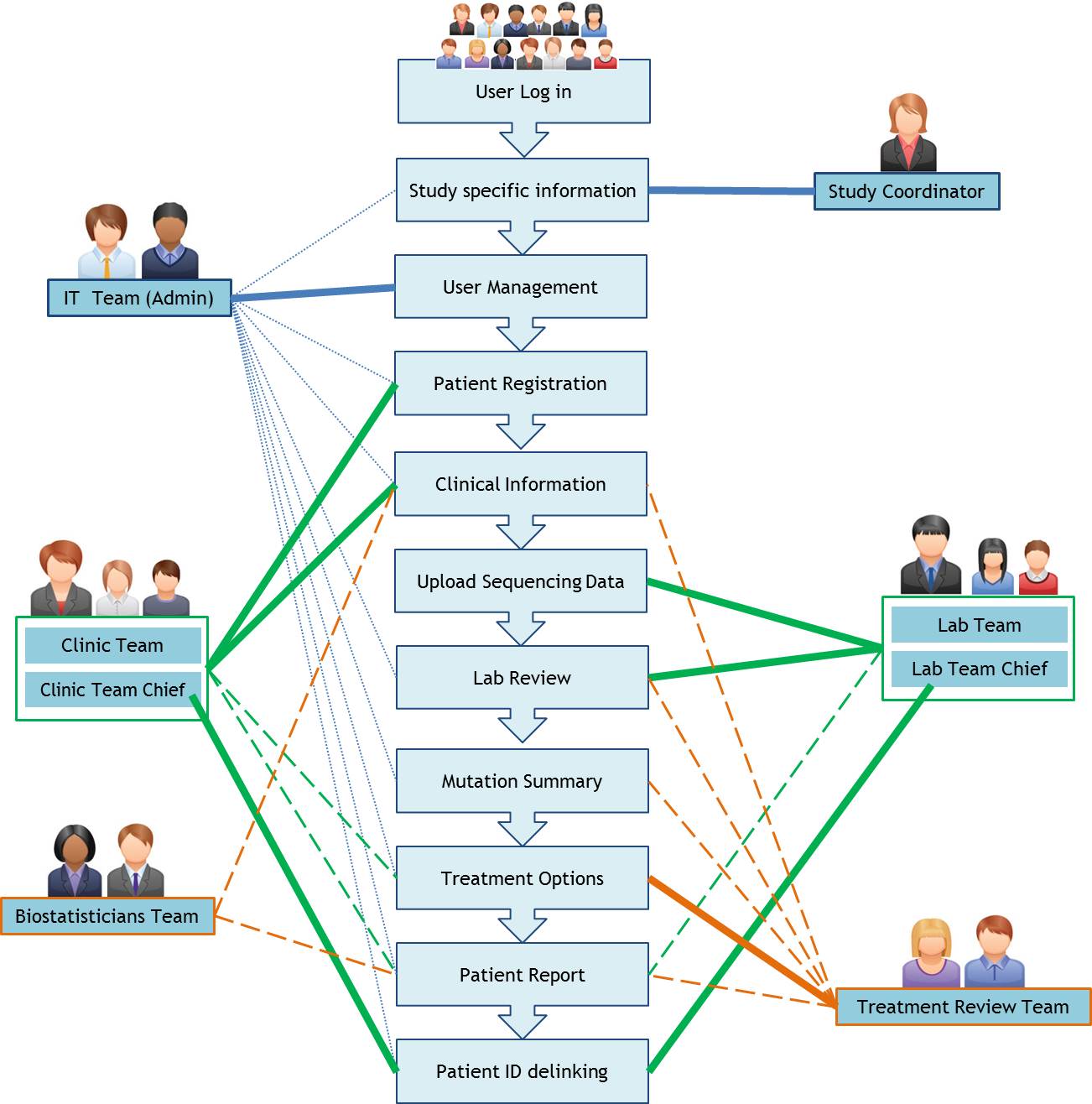
A typical workflow
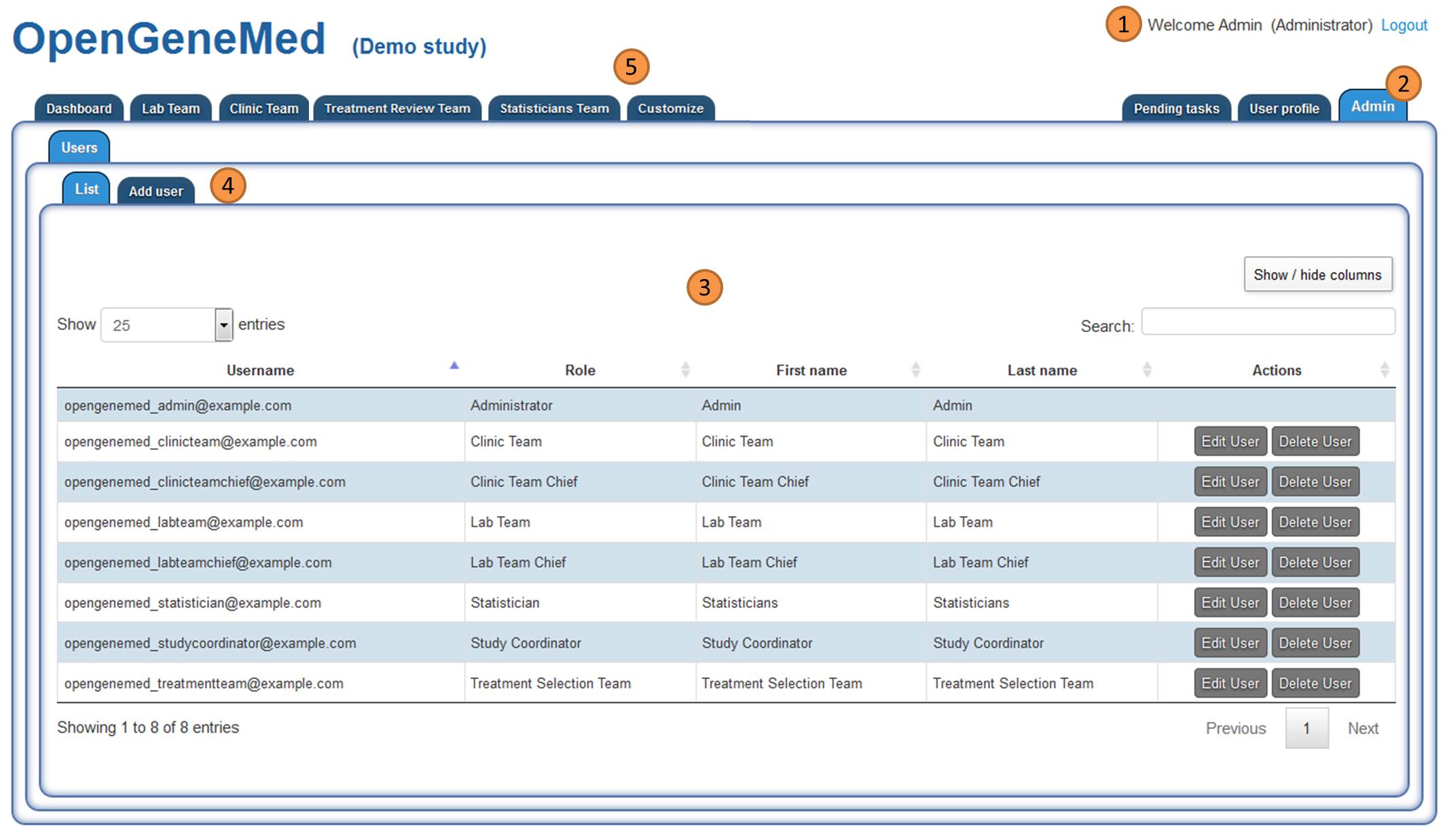
Admin tab
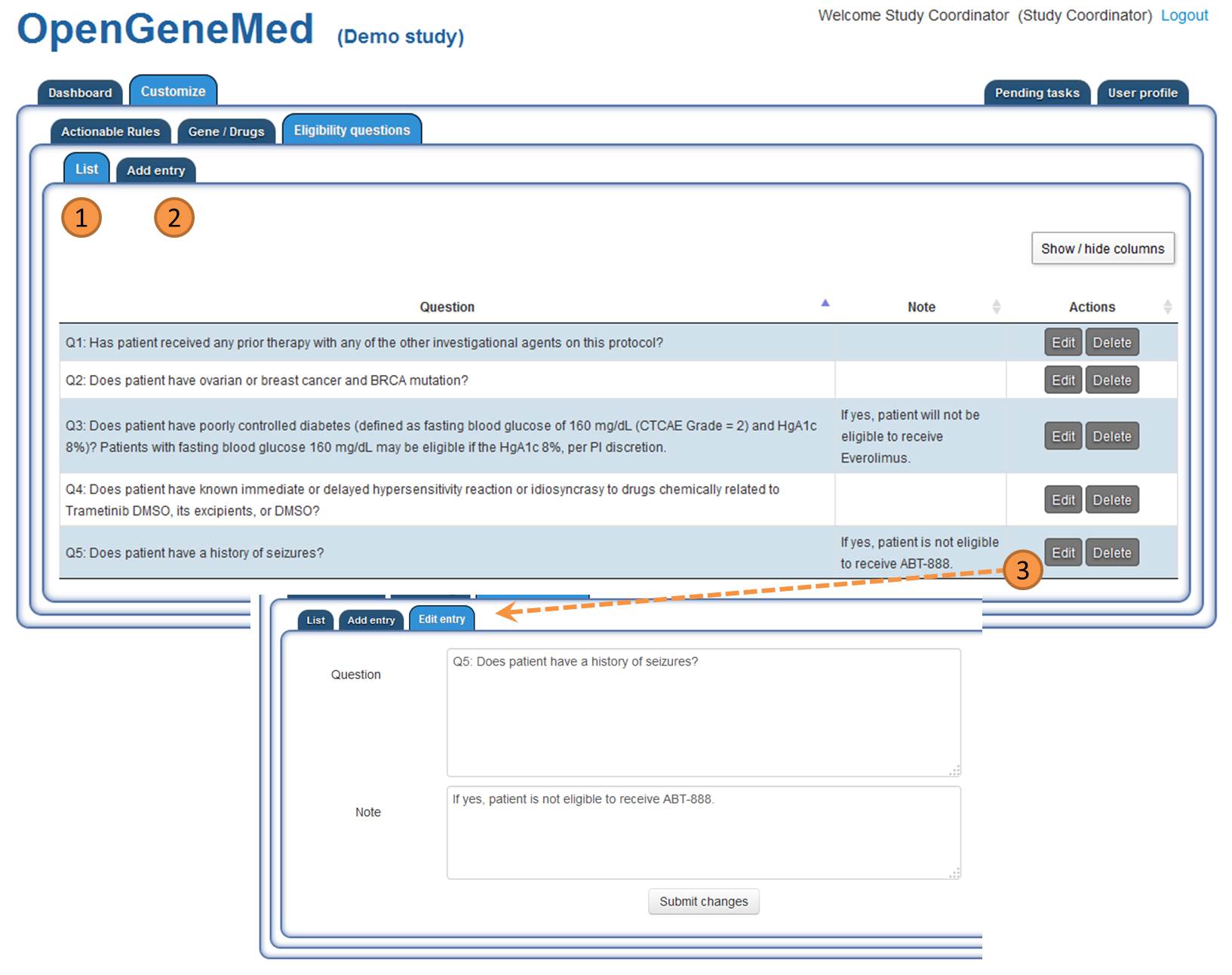
Study coordinator: customizable eligibility questions
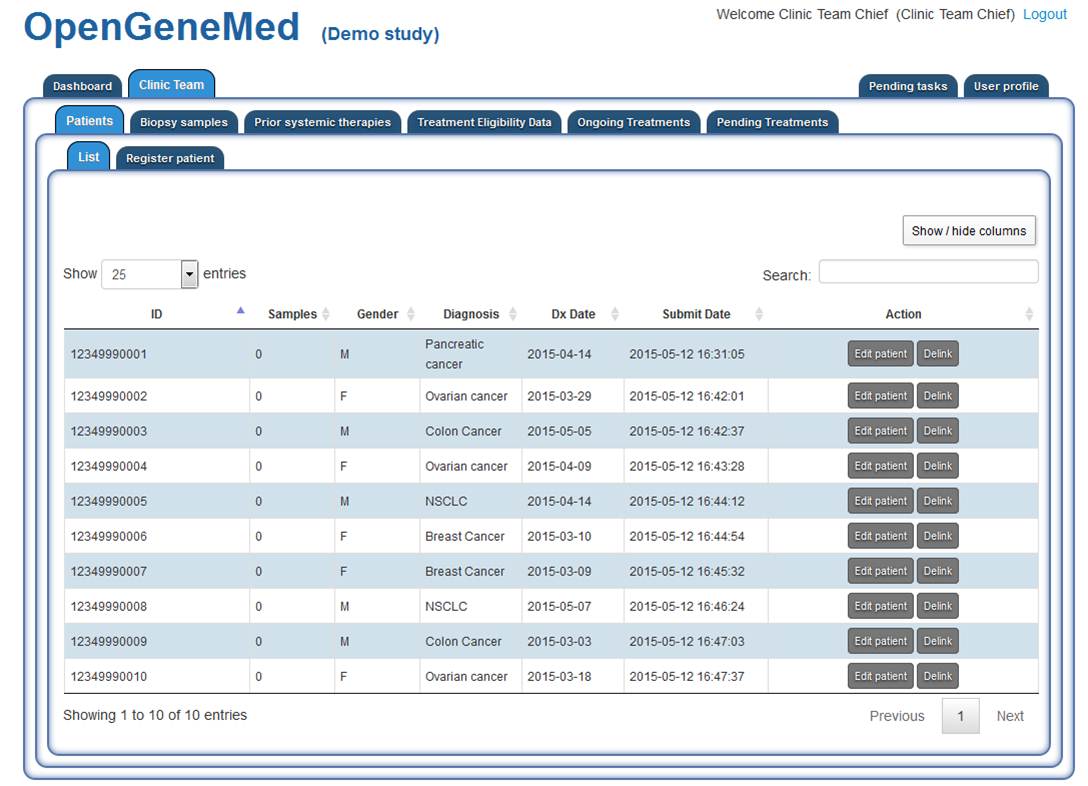
Clinic team: patient overview
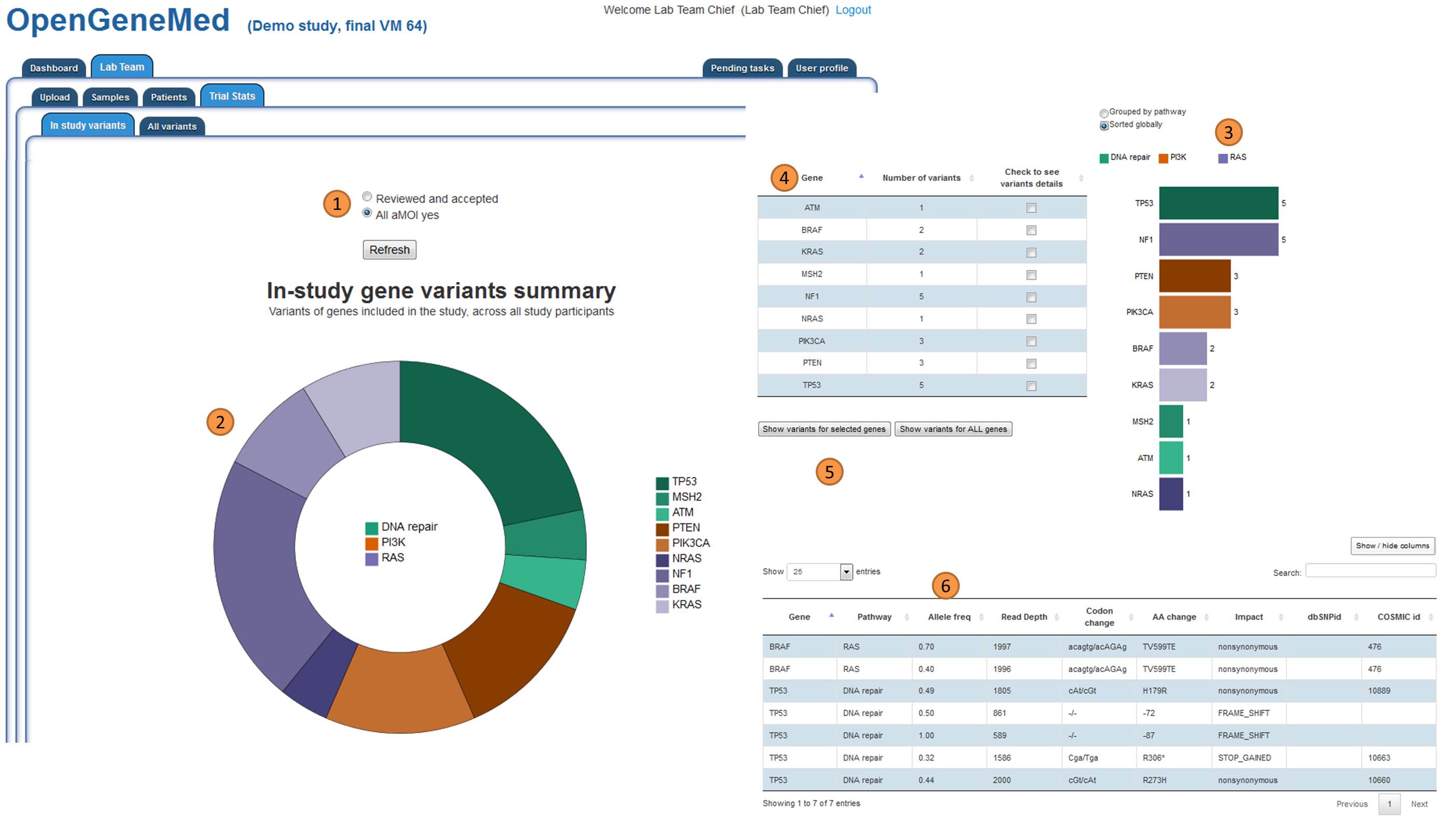
Biostatistician Team: study statistics
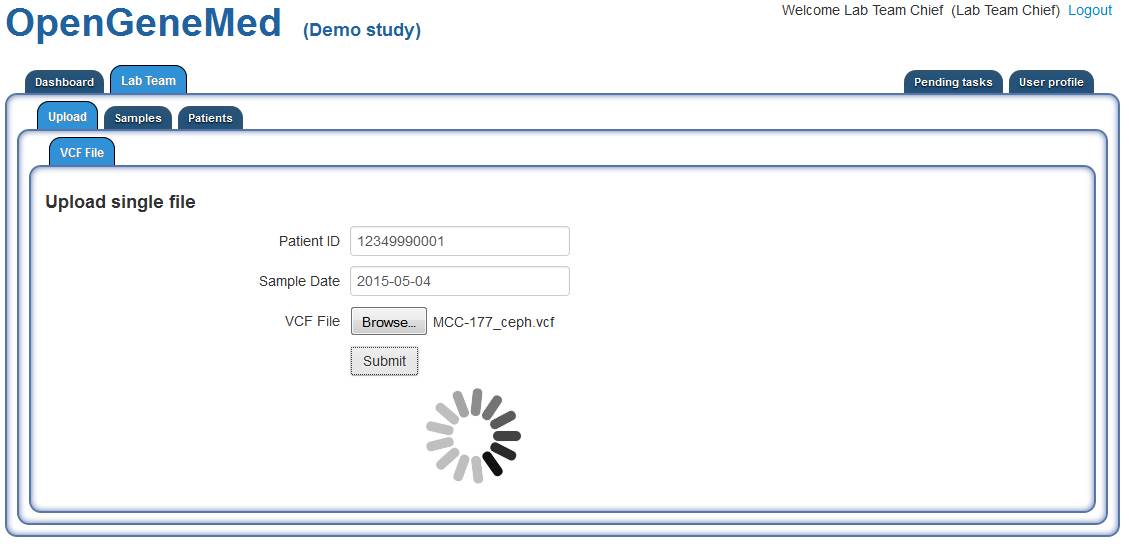
Upload VCF for annotation
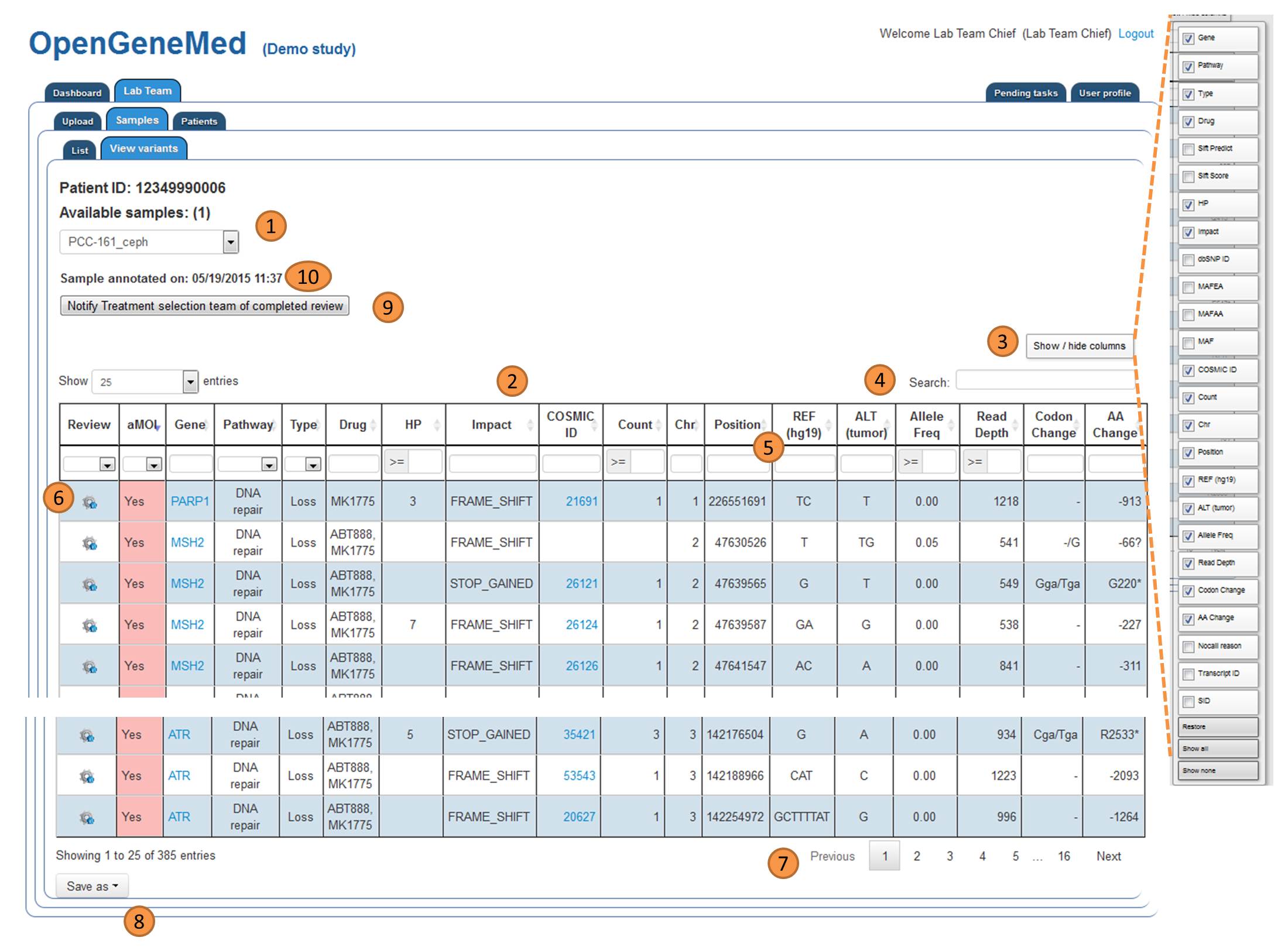
Table with annotation information





